Identification
CAS Number
19171-19-8
Name
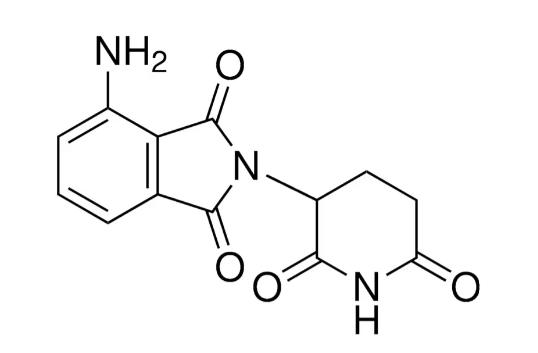
Pomalidomide
Synonyms
1,3-dioxo-2-(2,6-dioxopiperidin-3-yl)-4-aminoisoindoline
19171-19-8 [RN]
1H-Isoindole-1,3(2H)-dione, 4-amino-2-(2,6-dioxo-3-piperidinyl)- [ACD/Index Name]
3-amino-N-(2,6-dioxo-3-piperidyl)phthalamide
4-Amino-2-(2,6-dioxo-3-piperidinyl)-1H-isoindol-1,3(2H)-dion [German] [ACD/IUPAC Name]
4-Amino-2-(2,6-dioxo-3-piperidinyl)-1H-isoindole-1,3(2H)-dione [ACD/IUPAC Name]
4-Amino-2-(2,6-dioxo-3-pipéridinyl)-1H-isoindole-1,3(2H)-dione [French] [ACD/IUPAC Name]
4-amino-2-(2,6-dioxopiperidin-3-yl)-2,3-dihydro-1H-isoindole-1,3-dione
4-amino-2-(2,6-dioxopiperidin-3-yl)isoindoline-1,3-dione
Actimid
D2UX06XLB5
MFCD12756407 [MDL number]
pomalidomida [Spanish] [INN]
pomalidomide [French] [INN]
Pomalidomide [INN] [USAN] [Wiki]
POMALIDOMIDE, (R)-
POMALIDOMIDE, (S)-
pomalidomidum [Latin] [INN]
Pomalyst [Trade name]
UNII :D2UX06XLB5
(R)-pomalidomide
[19171-19-8] [RN]
1377838-49-7 [RN]
1H-Indol-7-ylmethanol [ACD/IUPAC Name]
202271-89-4 [RN]
202271-90-7 [RN]
2EE4M42K6G
3-Amino-N-(2,6-dioxo-3-piperidyl)phthalimide
443912-23-0 [RN]
443919-33-3 [RN]
4-amino-2-(2,6-dioxo-3-piperidinyl)isoindole-1,3-dione
4-Amino-2-(2,6-dioxo-3-piperidyl) isoindoline -1,3-dione
4-amino-2-(2,6-dioxo-3-piperidyl)isoindole-1,3-dione
4-Amino-2-(2,6-dioxo-3-piperidyl)isoindoline-1,3-dione
4-amino-2-(2,6-dioxopiperidin-3-yl)-1H-isoindole-1,3(2H)-dione
4-amino-2-(2,6-dioxopiperidin-3-yl)isoindole-1,3-dione
4-Amino-2-(2,6-dioxo-piperidin-3-yl)-isoindole-1,3-dione
4-AMINO-2-(2,6-DIOXO-PIPERIDIN-3-YL)ISOINDOLE-1,3-DIONE
4-Aminothalidomide
Actimid,4-Aminothalidomide,CC 4047
CC4047
CC-4047;Actimid ;CC4047 ;CC 4047
CC-4047|IMID-3|Imnovid®|Pomalyst®
D08976
DB08910
IMID-3
Imnovid
Imnovid (trade name)
Imnovid®
NR3397905
Phthalimide, 3-amino-N-(2,6-dioxo-3-piperidyl)-
pomalidomida
Pomalidomide (CC-4047)
Pomalidomide (JAN/USAN/INN)
pomalidomide(cc-4047,actimid)
Pomalidomide-d5
pomalidomidum
Pomalyst (TN)
Pomalyst®
помалидомид [Russian]
بوماليدوميد [Arabic]
泊马度胺 [Chinese]
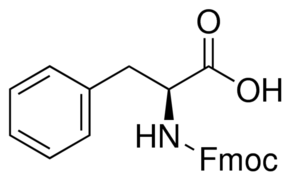
SMILES
c1cc2c(c(c1)N)C(=O)N(C2=O)C3CCC(=O)NC3=O
StdInChI
InChI=1S/C13H11N3O4/c14-7-3-1-2-6-10(7)13(20)16(12(6)19)8-4-5-9(17)15-11(8)18/h1-3,8H,4-5,14H2,(H,15,17,18)
StdInChIKey
UVSMNLNDYGZFPF-UHFFFAOYSA-N
Molecular Formula
C13H11N3O4
Molecular Weight
273.24
MDL Number
MFCD12756407
Properties
Appearance
White powder
Safety Data
RIDADR
NONH for all modes of transport
Specifications and Other Information of Our Pomalidomide CAS 19171-19-8
Identification Methods
HNMR, HPLC
Purity
95% min
Storage
Store at 2-8℃ for long time.
Features
Substrate Proteins : The primary ligands for E3 ligases are substrate proteins that need to be ubiquitinated for degradation or regulation. These substrate proteins can vary widely depending on the specific E3 ligase and the cellular process it regulates.
Ubiquitin : In the ubiquitination process, ubiquitin itself acts as a ligand. It forms a thioester bond with the active site cysteine of the E3 ligase before being transferred to the substrate protein.
Adaptor Proteins : Some E3 ligases require adaptor proteins to facilitate substrate recognition and ubiquitination. These adaptor proteins can also act as ligands for the E3 ligase.
Small Molecules : In some cases, small molecules or chemical compounds can modulate the activity of E3 ligases by binding to them directly or affecting their interactions with other proteins.
Post-translational Modifications (PTMs): PTMs of either the E3 ligase itself or its substrate proteins can also serve as ligands, regulating the ubiquitination process.
Overall, the ligands for E3 ligases play crucial roles in substrate recognition, ubiquitin transfer, and the regulation of cellular processes.
Known Application
Substrate Recognition : Ligands facilitate the recognition of specific substrate proteins by E3 ligases. These ligands can be protein motifs, post-translational modifications (such as phosphorylation or acetylation), or adaptor proteins that bring substrates to the E3 ligase.
Ubiquitination : Once a substrate is recognized, E3 ligases catalyze the transfer of ubiquitin molecules to the substrate protein. These ligands are crucial for facilitating the transfer of ubiquitin from the E2 ubiquitin-conjugating enzyme to the substrate.
Regulation of Protein Stability : Ubiquitination mediated by E3 ligases marks substrate proteins for degradation by the proteasome. Ligands for E3 ligases thus regulate the stability of proteins in the cell by targeting them for degradation.
Regulation of Protein Function : Ubiquitination can also regulate the activity, localization, or interaction partners of substrate proteins without leading to degradation. Ligands for E3 ligases are involved in this regulatory process by controlling the extent and site of ubiquitination.
Cellular Signaling : Ubiquitination mediated by E3 ligases is involved in various cellular signaling pathways, including those regulating cell cycle progression, DNA repair, immune response, and apoptosis. Ligands for E3 ligases modulate these signaling pathways by targeting specific proteins for ubiquitination.
Overall, ligands for E3 ligases are essential for substrate recognition, ubiquitination, and the regulation of protein stability and function, thereby influencing numerous cellular processes and signaling pathways.
Links
This product is developed by our R&D company Caming Pharmaceutical Ltd (https://www.caming.com/).
Quick Inquiry
Fill out our inquiry form and one of our experts will be in touch with you shortly.